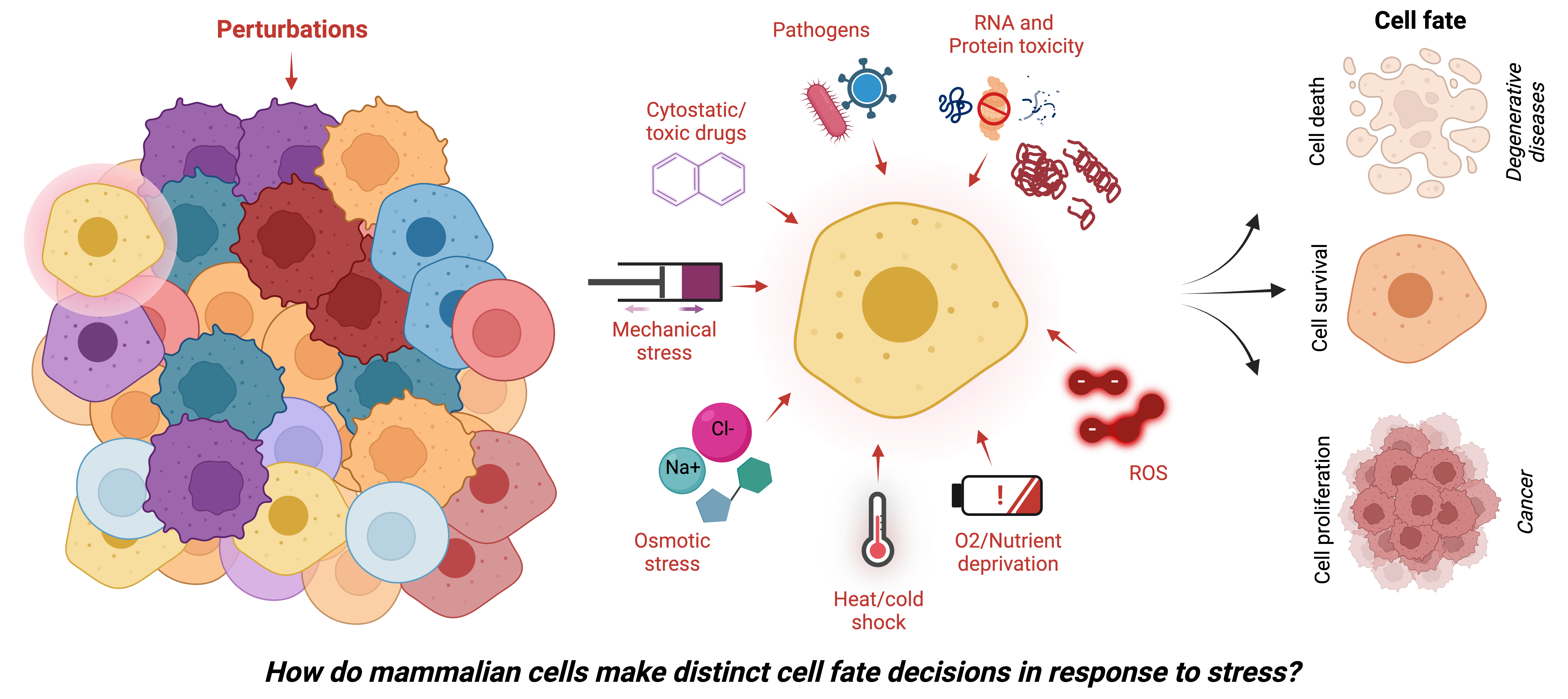
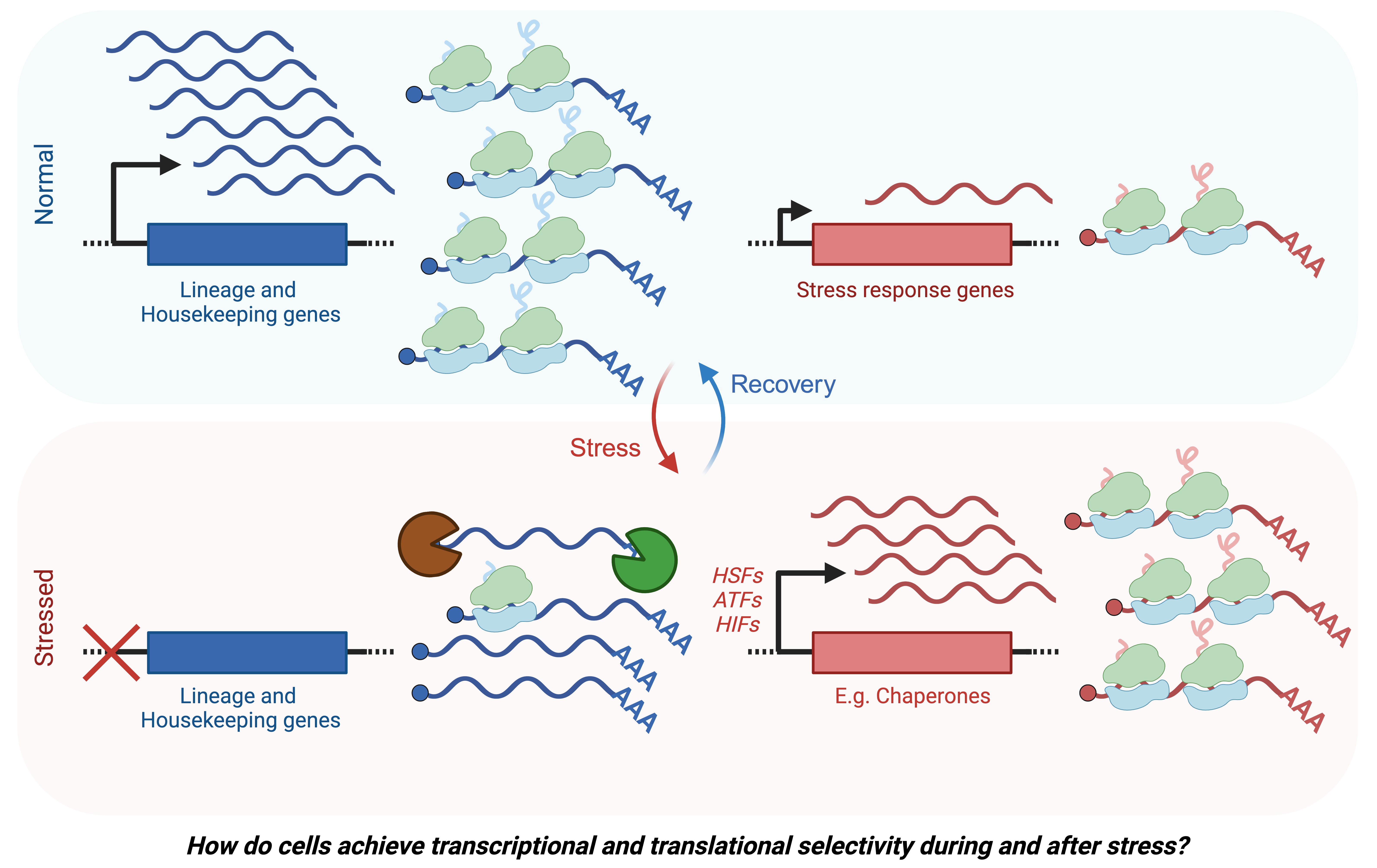
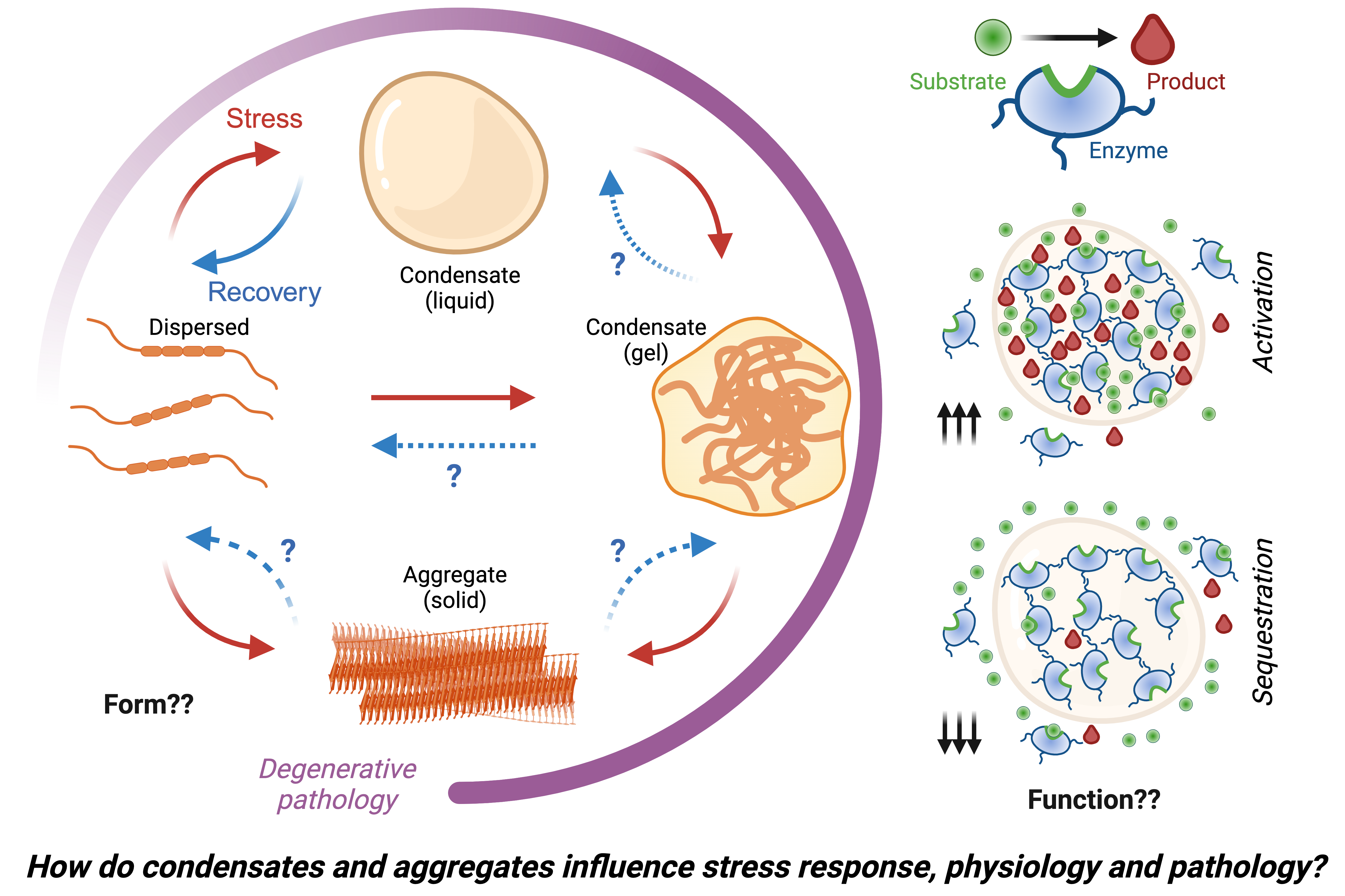
How do cells deal with stress?
Gene regulation || Biomolecular condensates || Cancer biology || Degenerative diseases || Single-molecule imaging || Single-cell and spatial omics
What we do?
We study how cell signaling, gene expression and subcellular organization influence cellular homeostasis and cell fate in health and disease.


Foundational Mechanisms
Diagnostics and Therapeutics
Technology Development
We investigate molecular mechanisms by which normal and malignant mammalian cells sense and respond to intrinsic and environmental stresses.
We test the diagnostic potential and therapeutic tractability of stress-responsive factors that influence cell survival in degenerative pathologies and cancer.
We develop imaging and sequencing technologies to spatiotemporally probe biomolecular organization and gene expression at high-sensitivity, -resolution and -throughput.
Our philosophy
Multifaceted perspectives lead to better science — and a better world.
Our mantra
Be bold, be ready, educate and enjoy.
Interested in joining our team?